Search and filtering¶
Search operations in CVA intend to support different use cases from computational analysis to front end features. For computational analysis we need a comprehensive filtering including some normalisation over text fields, arithmetic operations (e.g.: count_samples > 3) and boolean operations (e.g.: this eq 1 or that eq 2 and this eq 1 and that eq 2). To support a front end we will also require some text search performing natural language processing (e.g.: metabolic matches metabolism and connector words are ignored), regex queries and ad hoc summaries to support autocomplete features.
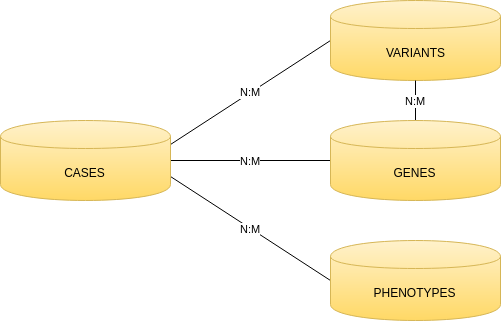
The principal entities that are available for search are: cases and report events. There are other secondary entities: variants, genes and phenotypes.
%run initialise_pyark.py
Filtering¶
Main entities has a main get endpoint that exposes a comprehensive set of filters. Some fields are normalised in the database and every query is normalised to achieve a case insensitive matching.
The normalised fields are:
- Panel names
- Disorders
- Interpretation service
Filters are combined with an AND operator¶
# get the first page of cases
cases = next(cases_client.get_cases(
program=Program.rare_disease,
assembly=Assembly.GRCh38,
specificDiseases='intellectual disability',
as_data_frame=True))
# same filters can be applied to count operations
cases_client.count(
program=Program.rare_disease,
assembly=Assembly.GRCh37,
specificDiseases='intellectual disability')
Normalisation allows us to make filters case insensitive¶
The filters on which normalisation is applied are:
- Panel names
- Disease names, groups, subgroups, etc.
- Interpretation service names
cases_client.count(specificDiseases='intellectual disability')
cases_client.count(specificDiseases='INTELLECTUAL DISABILITY')
Cross references¶
Phenotypes and genes are stored as identifiers in CVA (ie: Ensembl identifiers for genes, HPO terms identifiers for rare disease phenotypes and SNOMED/ICD10 for cancer phenotypes) but they can be matched by cross references to other databases.
We can search genes by ensembl identifier, HGNC gene symbol, gene synonym, Uniprot, CCDS and LRG.
http://www.ensembl.org/Homo_sapiens/Gene/Summary?db=core;g=ENSG00000130164;r=19:11089362-11133816
NOTE: these queries call CellBase and thus have a lower performance
# match by gene Ensembl identifier
cases_client.count(genes='ENSG00000130164')
# match by HGNC gene symbol
cases_client.count(genesXrefs='LDLR')
# match by synonym
cases_client.count(genesXrefs='LDLCQ2')
# match by Uniprot identifier
cases_client.count(genesXrefs='P01130')
# match by Human CCDS
cases_client.count(genesXrefs='CCDS12254.1')
# match by LRG region
cases_client.count(genesXrefs='LRG_274')
We can search HPO terms by identifier, alternative HPO id, SNOMED-CT (US version), UMLS and MSH.
https://hpo.jax.org/app/browse/term/HP:0003701
NOTE: alternative HPO terms are replaced automatically by the most current HPO term when queried
cases_client.count(phenotypes='HP:0003701')
# search by UMLS
cases_client.count(phenotypesXrefs='UMLS:C0221629')
# search by SNOMED CT
cases_client.count(phenotypesXrefs='SNOMEDCT_US:249939004')
# search by alternative HPO id
cases_client.count(phenotypesXrefs='HP:0008961')
Boolean filtering¶
By default filtering results match all filters, as opposed to match any filter. There are two workarounds for this behaviour:
- The
filterparameter - The text search (see section below)
The filter parameter follows a basic subset of the OData specification for $field https://www.odata.org/documentation/odata-version-3-0/url-conventions/
Following this specification we define equality and inequality with the operators eq and ne, multiple expressions can be combined with or and and operators.
There are known limitations to this filtering approach:
- there is no normalisation happening in query values
- the field names have to follow the schema naming in the database which is very obscure to the user
- we are not able to combine different boolean operators (ie:
(this and that) or those) - we do not support substring operations
# AND operator
cases_client.count(
filter="pedigreeAnalysisPanels.specificDisease eq 'intellectual disability' and " +
"probandDisorders.specificDisease eq 'intellectual disability'")
# OR operator
cases_client.count(
filter="pedigreeAnalysisPanels.specificDisease eq 'intellectual disability' or " +
"probandDisorders.specificDisease eq 'intellectual disability'")
# AND operator and negation
cases_client.count(
filter="pedigreeAnalysisPanels.specificDisease eq 'intellectual disability' and " +
"probandDisorders.specificDisease ne 'intellectual disability'")
# normalisation is not applied
cases_client.count(
filter="pedigreeAnalysisPanels.specificDisease eq 'INTELLECTUAL DISABILITY' and " +
"probandDisorders.specificDisease ne 'intellectual disability'")
Also when provided a list of values the default behaviour is that results match any element in the list. There is a workaround for this behaviour in a specific use case: phenotypes.
# default behaviour for a list of phenotypes
cases_client.count(phenotypes=["HP:0005338", "HP:0002000"])
# explicitly matching any phenotype
cases_client.count(phenotypes=["HP:0005338", "HP:0002000"], anyOrAllPhenotypes='ANY')
# matching all phenotypes
cases_client.count(phenotypes=["HP:0005338", "HP:0002000"], anyOrAllPhenotypes='ALL')
# also works for cross references
cases_client.count(phenotypesXrefs=["HP:0005338", "UMLS:C1857479"], anyOrAllPhenotypes='ALL')
Arithmetic filtering¶
The filter parameter also support arithmetic operations: lt, le, gt and ge on numeric values.
There is no other alternative to perform arithmetic comparisons than with the filter parameter.
# search for cases with at least 3 samples and the proband being born after 2016
cases_client.count(filter="countSamples ge 3 and probandYearOfBirth gt 2016")
cases_client.count(
filter="pedigreeAnalysisPanels.specificDisease eq 'intellectual disability' and " +
"countTiers.TIER1 eq 0 and countTiers.TIER2 eq 0")
Search¶
The previous filtering approaches are fit for most use cases specially for a computational analysis, but to support human friendly search operations from a front end we need some natural language processing to do partial matching, tokenisation, stemming and ranking of matches. The results from a text search will be fuzzier and not exact as compared to results provided by filtering.
There are two main use cases:
- Free text search. A word or a set of words give a number of documents
- Autocomplete. A partial word gives a number of possible matches for that word across all existing documents
Free text search¶
There are two entities where search is supported: cases, phenotypes and variants. This is available through /cases?search=something, /variants?search=something and /hpos/search?search=something
The search of cases supports the following:
- Identifier based search
- Case identifier (either unversioned "12345" or versioned "12345-2")
- Family identifier
- Participant identifier (ie: any participant in the family, not only the proband)
- HPO identifier (ie: HPOs present in the proband)
- Gene id + gene cross references (ie: genes having a variant in the case)
- Variant cross references (ie: variants in the case)
- Semantic search
- Clinical indication
- Panel name
- HPO name
- HPO synonyms
- Other regex based search
- Variant coordinates (ie: cases with this variant)
- Genomic region (ie: cases with any variant within this region)
The search of variants supports the following:
- Identifier based search
- Variant cross references (ie: ClinVar, dbSNP, COSMIC)
- Gene cross references (ie: Ensembl gene and transcript, Uniprot accession, name and variant id, HGNC gene symbol)
- Other "things" related with the gene (ie: PubMed id, gene drug interactions (e.g.: DGIdb))
- Other regex based search
- Variant coordinates
- Genomic region
The search of phenotypes supports the following:
- Semantic search
- Name
- Synonyms
# exact match of metabolic against disorder or panel name
cases_client.count(search="metabolic")
# match of stem of the word against disorder or panel name
cases_client.count(search="metabolism")
# stop words are ignored
cases_client.count(search="metabolism of this and that")
# match "metabolism" or "undiagnosed"
cases_client.count(search="metabolic undiagnosed")
# match exactly "metabolism" and "undiagnosed" by quaoting each word independently
cases_client.count(search='"metabolic" "undiagnosed"')
# match an exact ordered set of words by quoting all words
cases_client.count(search='"undiagnosed metabolic disorders"')
# numeric values will match any numeric field containing it
cases_client.count(search='10000-1')
# search by variant
cases_client.count(search='8:8377111:A:T')
# search by gene
cases_client.count(search='ENSG00000213516')
# search by gene
cases_client.count(search='BRCA1')
HPO terms can be searched over the canonical name, the definition and the synonyms.
hpos = next(entities_client.get_hpos(search='"color blind"', as_data_frame=True))
hpos[["identifier", "name", "synonyms"]]
Autocomplete search¶
Cases have a dedicated endpoint to support autocomplete functionality based on panels and disorders which is in /cases/search.
The response contains scored matches of different categories which will potentially allow to suggest to the user a sorted set of named things.
cases_client.search('metabol')
Genes and their cross references can be matched by prefix.
# match cross reference
entities_client.get_genes(xrefRegex="^BRCA", as_data_frame=True).geneSymbol
# match gene symbols
entities_client.get_genes(geneSymbolRegex="^BRCA", as_data_frame=True).geneSymbol
Panels can be matched by regex.
entities_client.get_panels_by_regex(regex="intellect", as_data_frame=True).head()
Disorders can be matched by regex.
entities_client.get_disorders_by_regex(regex="intellect", as_data_frame=True).head()
HPO terms and their cross references can be matched by regex.
next(entities_client.get_hpos(xrefRegex="HP:000001", as_data_frame=True)).identifier
next(entities_client.get_hpos(xrefRegex="SNOMED.*123", as_data_frame=True)).identifier